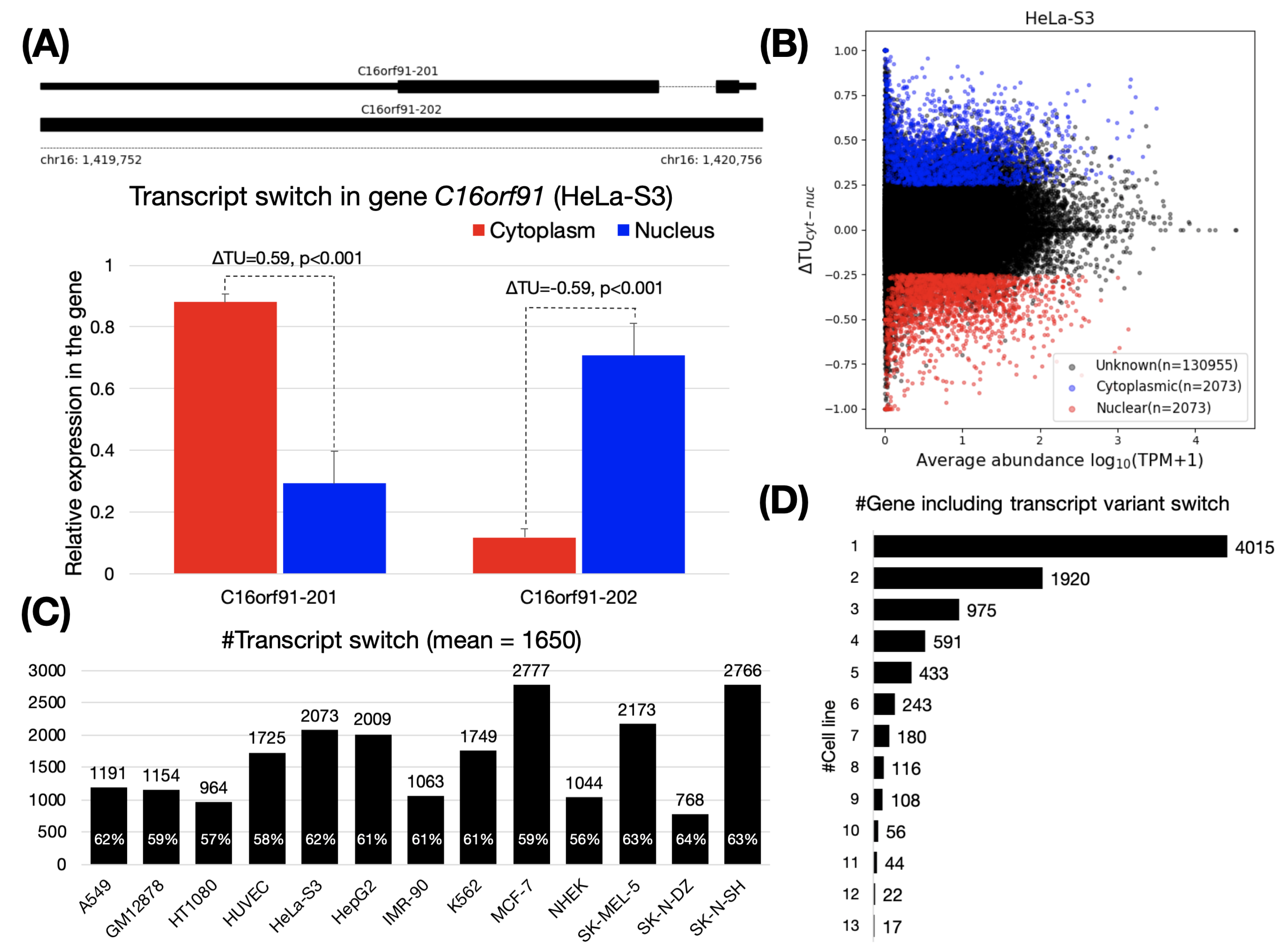
Quantitative visualization of alternative exon expression from RNA-Seq data - Analysis of RNA sequencing (RNA-Seq)… | Visualisation, Data visualization, Expressions

Alternative splicing and splice isoform analysis with Iso-Seq reads.... | Download Scientific Diagram
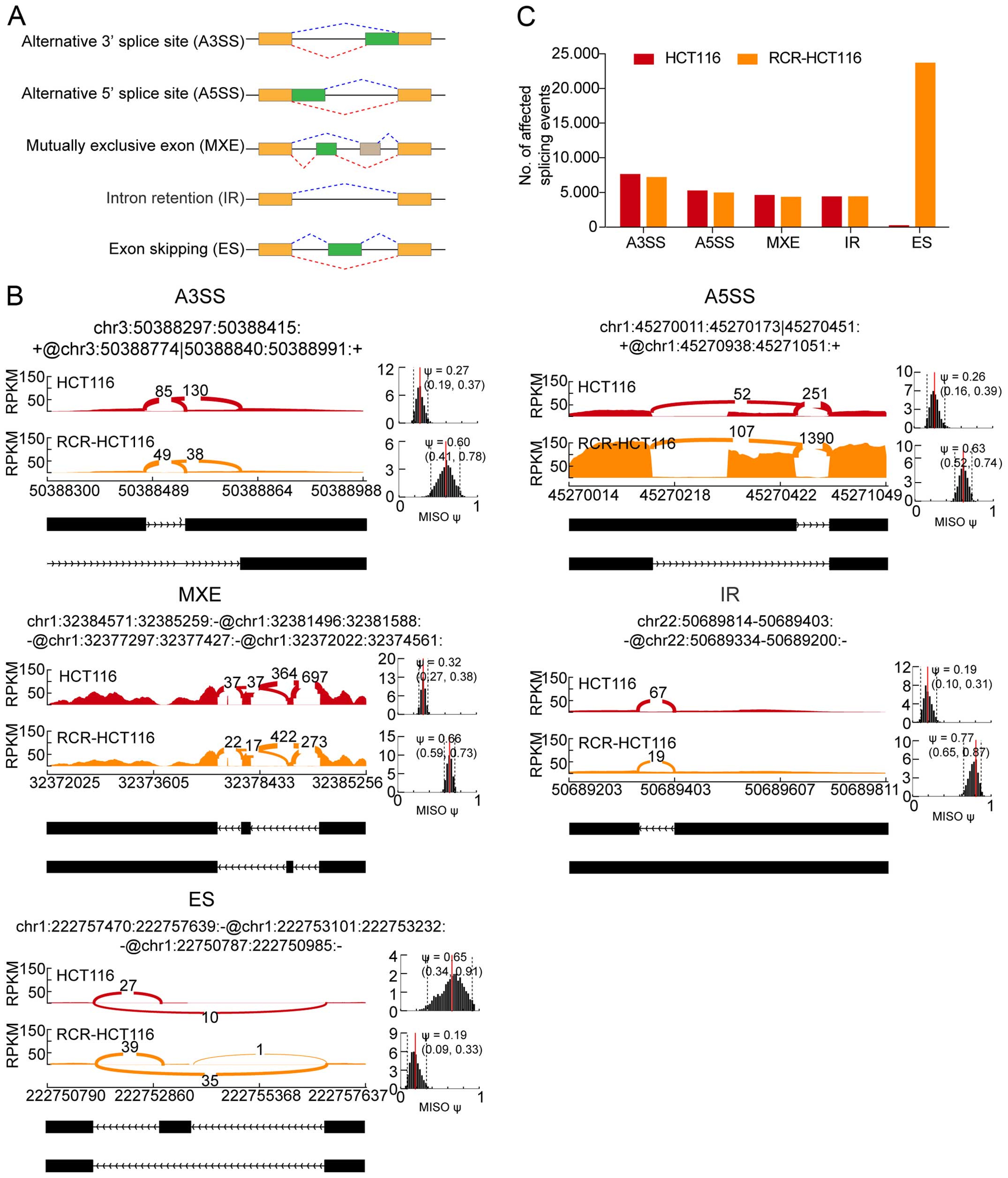
Genome-wide profiling of chemoradiation‑induced changes in alternative splicing in colon cancer cells

EBChangePoint – An empirical Bayes change-point model for identifying 3′ and 5′ alternative splicing by RNA-Seq | RNA-Seq Blog
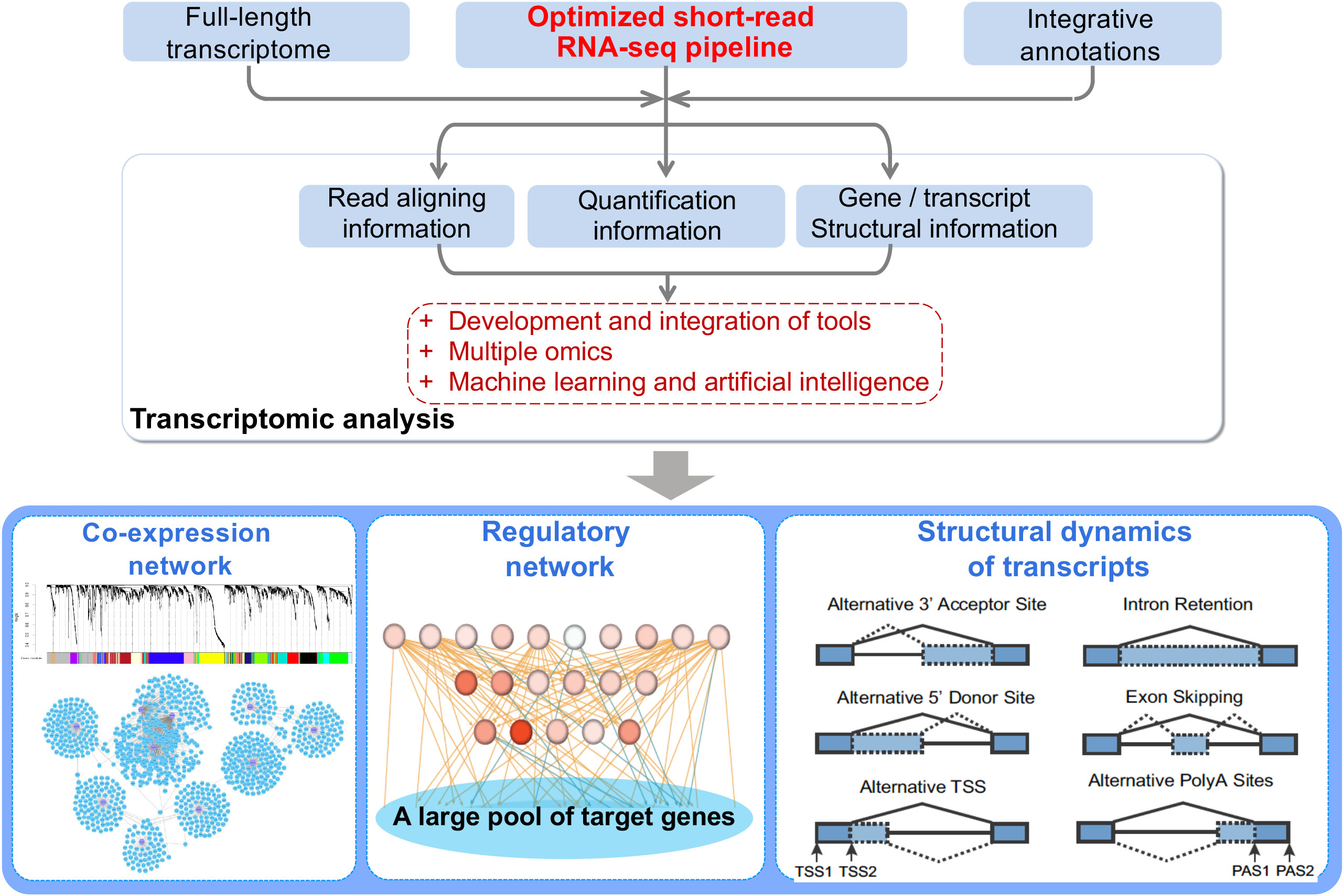
Frontiers | Unleashing the power within short-read RNA-seq for plant research: Beyond differential expression analysis and toward regulomics
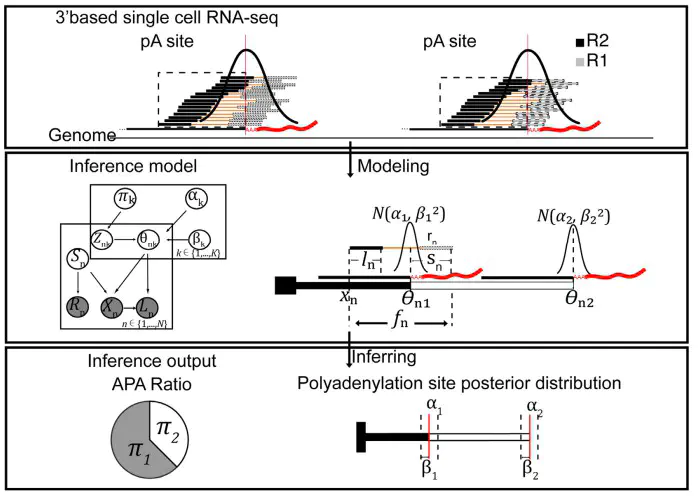
Our paper about estimating alternative polyadenylation (APA) from scRNA-seq has been accepted by Nucleic Acids Research. | Aalto Computational Genomics Lab
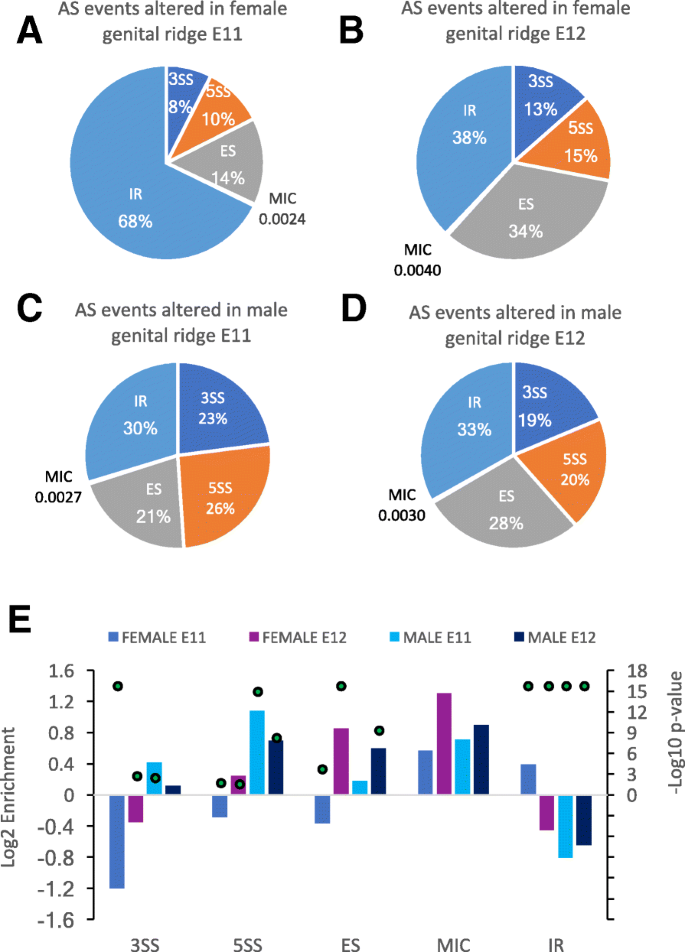
Differential isoform expression and alternative splicing in sex determination in mice | BMC Genomics | Full Text
![PDF] Detection, annotation and visualization of alternative splicing from RNA-Seq data with SplicingViewer. | Semantic Scholar PDF] Detection, annotation and visualization of alternative splicing from RNA-Seq data with SplicingViewer. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d09ff8a3b1debf3f4a38d5ae620cdbc855b9962f/2-Figure1-1.png)
PDF] Detection, annotation and visualization of alternative splicing from RNA-Seq data with SplicingViewer. | Semantic Scholar
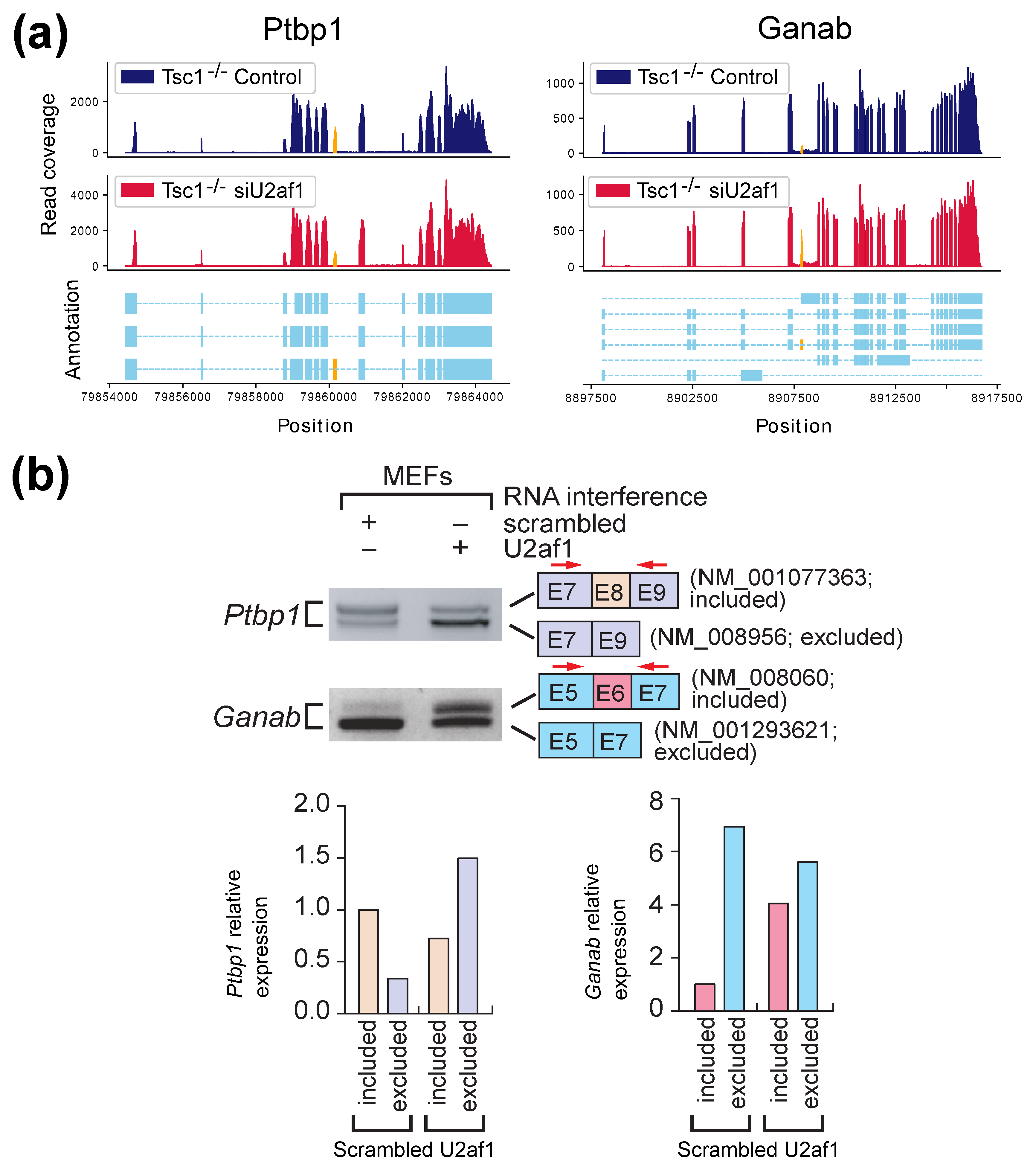
IJMS | Free Full-Text | AS-Quant: Detection and Visualization of Alternative Splicing Events with RNA-seq Data

Analysis of Multiple Years of RNA-Seq Data Using the 3D-RNA-Seq Application Reveals Seasonal Signatures of Differential Gene and Transcript-Level Expression, Alternative-Splicing, and Transcript... — Journal of Young Investigators